 |
 |
| Anim Biosci > Volume 29(3); 2016 > Article |
|
Abstract
Lipins play dual function in lipid metabolism by serving as phosphatidate phosphatase and transcriptional co-regulators of gene expression. Mammalian lipin proteins consist of lipin1, lipin2, and lipin3 and are encoded by their respective genes Lpin1, Lpin2, and Lpin3. To date, most studies are concerned with Lpin1, only a few have addressed Lpin2 and Lpin3. Ontogenetic expression of Lpin2 and Lpin3 and their associations with traits would help to explore their molecular and physiological functions in sheep. In this study, 48 animals with an equal number of males and females each for both breeds of fat-tailed sheep such as Guangling Large Tailed (GLT) and Small Tailed Han (STH) were chosen to evaluate the ontogenetic expression of Lpin2 and Lpin3 from eight different tissues and months of age by quantitative real-time polymerase chain reaction (PCR). Associations between gene expression and slaughter and tail traits were also analyzed. The results showed that Lpin2 mRNA was highly expressed in perirenal and tail fats, and was also substantially expressed in liver, kidney, reproductive organs (testis and ovary), with the lowest levels in small intestine and femoral biceps. Lpin3 mRNA was prominently expressed in liver and small intestine, and was also expressed at high levels in kidney, perirenal and tail fats as well as reproductive organs (testis and ovary), with the lowest level in femoral biceps. Global expression of Lpin2 and Lpin3 in GLT both were significantly higher than those in STH. Spatiotemporal expression showed that the highest levels of Lpin2 expression occurred at 10 months of age in two breeds of sheep, with the lowest expression at 2 months of age in STH and at 8 months of age in GLT. The greatest levels of Lpin3 expression occurred at 4 months of age in STH and at 10 months of age in GLT, with the lowest expression at 12 months of age in STH and at 8 months of age in GLT. Breed and age significantly influenced the tissue expression patterns of Lpin2 and Lpin3, respectively, and sex significantly influenced the spatiotemporal expression patterns of Lpin3. Meanwhile, Lpin2 and Lpin3 mRNA expression both showed significant correlations with slaughter and tail traits, and the associations appear to be related with the ontogenetic expression as well as the potential functions of lipin2 and lipin3 in sheep.
Lipin proteins act as Mg2+-dependent phosphatidate phosphatase (PAP) and catalyze the penultimate step of converting phosphatidic acid (PA) to diacylglycerol (DAG) in the glycerol phosphate pathway (Han et al., 2006; Donkor et al., 2007). Lipins also work as the transcriptional co-regulators of gene expression in the nucleus to promote fatty acid oxidation (Péterfy et al., 2010; Chen et al., 2012). Mammals express three lipin proteins, namely lipin1, lipin2 and lipin3, and are encoded by their respective genes Lpin1, Lpin2, and Lpin3 (Péterfy et al., 2001). Lipin function has been evolutionarily conserved from a single ortholog in yeast to the mammalian lipin family proteins because of the highly conserved functional domains (Péterfy et al., 2001).
Lipins play crucial roles in maintaining the balance of lipid intermediates such as PA and DAG (Dwyer et al., 2012; Mitra et al., 2013). Mutations in Lpin1 in fld (the fatty liver dystrophy) mice lead to lipodystrophy and neonatal fatty liver (Reue et al., 2000), and overexpression of Lpin1 in adipose tissue or skeletal muscle promotes adiposity (Reue et al., 2000; Phan and Reue, 2005). Mutations in human Lpin1 cause severe myoglobinuria in childhood (Michot et al., 2010), and those in Lpin2 are associated with an autoinflammatory bone disease known as Majeed syndrome (Herlin et al., 2013). A variant near Lpin3 has been associated with fasting serum glucose levels in healthy individuals (Scott et al., 2012).
Lipin proteins and their encoding genes exhibit distinct yet overlapping tissue expression patterns (Donkor et al., 2007). Mouse lipin proteins are all PAP activity and lipin1 has substantially higher PAP specific activity than lipin2 and lipin3 (Donkor et al., 2007). Lipin proteins appear to differ in molecular regulation and physiological function and can cooperate with each other in some physiological contexts (Sanderson et al., 2009; Dwyer et al., 2012). Moreover, lipin proteins are implicated in other functions such as reproductive biology (Huang et al., 2011; González et al., 2012) and maintaining nuclear/endoplasmic reticulum (ER) membrane morphology in mammalian or lower organisms (Golden et al., 2009; Gorjánácz and Mattaj, 2009; Sasser et al., 2012).
Lipins have been widely studied in mammals, however, most publications focus on lipin1, fewer of those have dealt with lipin2 and lipin3. In this study, we examined the ontogenetic expression of Lpin2 and Lpin3 in two breeds of fat-tailed sheep with phenotypic differences in tail fat phenotype, and analyzed the associations between gene expression and slaughter and tail traits, with the aims to reveal the ontogenetic expression patterns of the two genes and explore the potential functions of lipin2 and lipin3 in fat metabolism as well as other possible metabolic pathways in sheep.
Two breeds of Chinese fat-tailed sheep, i.e. Guangling Large Tailed (GLT) and Small Tailed Han (STH), both originated from the ancient Mongolia sheep, however, they varied in appearance and performance due to their geographic distribution and local acclimatization. GLT is characterized by large tail but low fecundity, STH is distinguished by high fecundity and a small tail. GLT is mainly distributed in the mountain regions of Shanxi province and STH in the plain regions of Shandong and Hebei provinces, China (Yuan et al., 2012).
Both breeds of lambs (1 month of age) were reared respectively in their local distribution regions during December. The feeding and management procedures were same, all the animals were raised in confinement during dry seasons in winter and spring while grazing on pasture in grass seasons. The dietary nutrition levels were established according to the Agricultural Standards (NY/T 816-2004) of the People’s Republic of China.
Four males and four females from every growth stage (2, 4, 6, 8, 10, and 12 months of age) totaling 48 animals for each breed were used to examine the ontogenetic expression of Lpin2 and Lpin3. Tissues such as liver, kidney, small intestine, femoral biceps, reproductive organs (testis and ovary), perirenal and tail fats were rapidly collected after slaughtering, weighed and placed in liquid nitrogen and stored at −80°C until subsequent laboratory analysis. Slaughter and tail traits such as tail length (cm), tail width (cm), body weight (kg), carcass weight (kg) and absolute tail fat weight were recorded, dressing percentage and relative tail fat weight were calculated, absolute tail fat weight (kg) was neglected in STH because of the small tail and low fat deposits. The feeding, management and slaughtering followed the National (GB13078–2001 and GB/T17237–1998) and the Agricultural Standards (NY5148-2002-NY 5151–2002) of the People’s Republic of China.
Total RNA was isolated using the TRIzol Reagent (Invitrogen, Carlsbad, CA, USA) and was purified with the RNeasy Mini Kit (QIAGEN GmbH, Hilden Germany) as recommended by the manufacturers’manual. Single-stranded cDNA was synthesized from one microliter of total RNA using the PrimeScript RT reagent Kit (Takara, Biotechnology, Dalian, China), according to the manufacturer’s guides. Target and housekeeping genes’ mRNA expression was carried out by Applied Biosystems 7500 Fast Real-Time PCR System (Applied Biosystems, Foster, CA, USA) and SYBR Premix Ex Taq II kit (Takara, Biotechnology, China) following the manufacturer’s instructions. Lpin2 and Lpin3 mRNA expression was normalized to housekeeping genes RPL13A and B2M mRNA levels respectively, data analysis was performed using the comparative CT method (Livak and Schmittgen, 2001), and the final relative expression was obtained by geometric mean. Primers (Table 1) were designed according to the sheep mRNA sequence (GenBank accession number XM_004020631 and XM_012155543.1) and those for sheep B2M and RPL13A by using online Primer3 plus (http://www.bioinformatics.nl/cgibin/primer3plus/primer3plus.cgi). PCR amplification procedure was as follows: 95°C for 30 s, 42 cycles at 95°C for 5 s and 61°C for 35 s, and a following cycle at 95°C for 15 s, 61°C for 1 min and 95°C for 15 s to obtain the dissociation curves. The standard curves were derived from a 2-fold dilution step by step with eight levels for target and housekeeping genes. PCR efficiencies of the four genes were between 90% and 110%. The R2 values of standard curves ranged from 0.974 to 0.998, and melting curves showed a single peak. Relative ratios of Lpin2 or Lpin3 to RPL13A and B2M calculated using the standard curves confirmed that the real-time PCR results were trustworthy. All the reactions were executed in triplicate.
The relative abundance of Lpin2 and Lpin3 mRNA was analyzed by using general linear model in SPSS Statistics 19.0 software package (SPSS Science, Chicago, IL, USA). Four factors including breed, sex, age, tissue and their two-way interactions were considered as the influencing factors on the mRNA expression. The model was fitted as the following function:
where, yijklm is the mRNA expression amount, μ the overall mean, Bi the ith breed (i = 1, 2), Sj the jth sex (j = 1, 2), Tk the kth tissue (k = 1, 2, ..., 8), Ml the lth months of age (l = 2, 4, ..., 12), BSij, BTik, BMil, STjk, SMjl, and TMkl correspond to the two-way interactions, and eijklm is the random residue. Multiple comparison among levels within different factors were performed by the Duncan’s method and the significance level was set at (p<0.05). The Pearson correlation coefficients between mRNA expression and traits were obtained by using SPSS Statistics 19.0 software package (SPSS Science, Chicago, IL, USA).
The relative abundance of Lpin2 and Lpin3 mRNA both displayed great differences in breeds (p<0.001, p<0.01), months of age (p<0.001) and tissues (p<0.001). Interactions between breed and age (p<0.001), breed and tissue (p<0.001, p<0.01), also tissue and age (p<0.05, p<0.01) significantly influenced Lpin2 and Lpin3 expression, respectively, interaction between sex and age (p<0.01) significantly influenced Lpin3 expression. Lpin2 and Lpin3 expression both showed no significant differences between males and females (p>0.05) (Table 2).
The average abundance of Lpin2 and Lpin3 mRNA for all tissues in GLT (1.042±0.042, 1.155±0.041) both were significantly higher than those in STH (0.671±0.043, 0.977±0.042) (p<0.001, p<0.01) (Table 3). Lpin2 expression in reproductive organs (testis and ovary) in GLT (0.876±0.067) was significantly higher than that in STH (0.487±0.068) (p<0.001), Lpin3 expression in reproductive organs (testis and ovary) showed no significant differences between GLT and STH (p>0.05). There were no significant differences of Lpin2 expression between testis and ovary (p>0.05), so did Lpin3 (data not shown).
Both Lpin2 and Lpin3 were expressed in all detected tissues, the highest level of Lpin2 expression was found in perirenal fat (1.587±0.079), with significant differences from those found in tail fat (p<0.05), liver, kidney and reproductive organs (testis and ovary) (p<0.01), as well as small intestine and femoral biceps (p<0.001). The greatest levels of Lpin3 expression were found in liver and small intestine (1.683±0.077, 1.539±0.077), respectively, with significant differences from those found in kidney (p<0.05), perirenal and tail fats, reproductive organs (testis and ovary) (p<0.01), as well as femoral biceps (p<0.001) (Table 3).
Lpin2 and Lpin3 mRNA abundance varied greatly along with the growth stages in both breeds of sheep, the greatest level of Lpin2 occurred at 10 months of age, which was much higher than those at 4, 12, and 2 months of age (p<0.05), with the lowest expression at 6 and 8 months of age (p<0.01). The maximum Lpin3 mRNA expression occurred at 4 and 10 months of age, respectively, which were much higher than those at 2, 12, and 6 months of age (p<0.05), with the lowest abundance at 8 months of age (p<0.01) (Table 3).
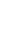
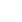
Two-way interactions significantly influenced Lpin2 mRNA expression. Figure 1 displays the interaction between breed and age, it suggested that the highest levels of Lpin2 mRNA occurred at 10 months of age in both breeds of sheep, however the lowest expression occurred at 2 months of age in STH and at 8 months of age in GTL, respectively. Figure 2 gives the effects of breed by tissue interaction on Lpin2 mRNA expression, it shows that Lpin2 was highly expressed in perirenal and tail fats in GLT, yet which was in tail fat and liver in STH, while the lowest levels were found in femoral biceps in both breeds of sheep. Figure 3 shows the change trends of tissue expression patterns along with age, the high expression of Lpin2 occurred sequentially in tail fat, perirenal fat and liver at 2 months of age, however, was found sequentially in perirenal fat, tail fat and liver at 4, 6, 10, and 12 months of age, and sequentially in perirenal fat, reproductive organs (testis and ovary) and tail fat at 8 months of age, while the lowest level was found in femoral biceps all through the growth stages.
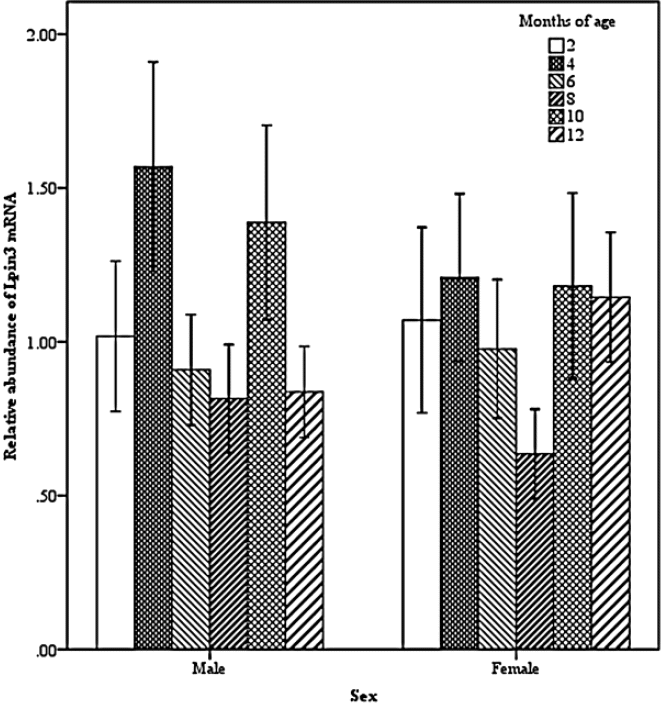
Two-way interactions also significantly impacted Lpin3 mRNA expression. Interaction between breed and age showed that the highest expression of Lpin3 occurred at 4 months of age in STH and at 10 months of age in GTL, with the lowest levels at 12 months of age in STH and at 8 months of age in GLT (Figure 4). Interaction between breed and tissue revealed that Lpin3 expression in perirenal and tail fats in GTL were higher than those in STH (Figure 5). Interaction between age and sex suggested that Lpin3 expression at 4 and 10 months of age in rams was higher than that in corresponding months of age in ewes (Figure 6). Interaction between tissue and age showed that the highest expression of Lpin3 was found in small intestine at 2, 4 months of age and yearlings, conversely, which was found in liver at 6, 8, and 10 months of age. The lowest expression was found in femoral biceps all through the growth stages except 8 months of age at which the lowest expression was found in reproductive organs (testis and ovary) (Figure 7).
Table 4 shows the associations between Lpin2 mRNA expression and slaughter and tail traits in both breeds of sheep. Associations analysis in GLT suggested that Lpin2 expression in liver was significantly negatively correlated with all the traits (p<0.01, p<0.05) but dressing percentage (p>0.05), and negative correlations were also found between the expression in femoral biceps and body weight, carcass weight (p<0.05). Lpin2 expression in reproductive organs (testis and ovary, esp. testis) was significantly positively correlated with all traits (p<0.01, p<0.05) but dressing percentage (p>0.05), however, those in perirenal and tail fats showed no significant correlations with traits in GLT (p>0.05) in spite of the high expression in these two tissues. In comparison, associations analysis in STH suggested that Lpin2 expression in liver, femoral biceps, perirenal and tail fats was significantly positively correlated with all the traits (p<0.01, p<0.05) but dressing percentage (p>0.05), and the positive correlations were also found between the expression in kidney and tail length (p<0.01), tail width and body weight (p<0.05).
Table 5 gives the associations between Lpin3 mRNA expression and slaughter and traits in both breeds of sheep. Association analysis in GLT showed that Lpin3 expression in liver, small intestine and tail fat was significantly positively correlated with the corresponding traits of absolute tail fat weight, dressing percentage and tail width (p<0.05), respectively, and the positive correlations were also found between the expression in perirenal fat and tail width and carcass weight (p<0.05), body weight and absolute tail fat weight (p<0.01). On the contrary, association analysis in STH showed that Lpin3 expression in liver was significantly negatively correlated with tail length, tail width and dressing percentage (p<0.05), and the negative correlations also occurred between the expression in small intestine and carcass weight (p<0.05) as well as dressing percentage (p<0.01).
Gene-expression studies in both mice and humans have demonstrated that Lpin2 is expressed at highest levels in liver and brain, with lower levels in the gastrointestinal tract (Donkor et al., 2007; Donkor et al., 2009). Lpin2 was detected in human adipose tissue but not in mouse adipose tissue (Donkor et al., 2007). The expression levels of Lpin3 are generally much lower than those of Lpin1 and Lpin2, with the highest levels in small intestine and bone (Donkor et al., 2009). Pig Lpin2 and Lpin3 are expressed highly in liver, and are also expressed at high levels in spleen, lung, kidney and adipose tissue, with low levels in heart and skeletal muscle, notably Lpin2 is high yet Lpin3 is low in pig small intestine (He et al., 2009). Chickens’ Lpin2 mRNA is liver and ovary enriched, with lowest levels in pancreas and leg muscles (Zhang et al., 2014). In our study, ovine Lpin2 was expressed at greatest levels in perirenal and tail fats, and was also substantially expressed in liver, kidney, reproductive organs (testis and ovary), with lowest expression in small intestine and femoral biceps. Ovine Lpin3 was expressed at maximum levels in liver and small intestine, and was also expressed highly in kidney, perirenal and tail fats as well as reproductive organs (testis and ovary), with low expression level in femoral biceps. The average expression of Lpin2 and Lpin3 in GLT both were higher than those in STH. Breed features show that GLT is typically of large tail and strong fat deposits, however, which is just opposite to STH (Yuan et al., 2012). Combined with the fat phenotype differences as well as the higher expression of the two genes in GLT, it could be concluded that Lpin2 and Lpin3 are the important regulators in fat-tailed sheep with active lipid metabolism.
Among three lipins, lipin1 is the dominant protein and accounts for the main PAP activity in adipose tissue and skeletal muscle cells (Donkor et al., 2007). Lipin1 expression increases during differentiation of the 3T3-L1 adipocyte cell line (Phan et al., 2004; Péterfy et al., 2005). Lipin2 can also be detected in preadipocytes but diminishes during differentiation (Grimsey et al., 2008). In addition, lipin1 deficiencies in humans are not associated with lipodystrophy as the fld mice (Michot et al., 2010), which might be compensated by the high expression of Lpin2 in adipose tissue (Donkor et al., 2007). Unlike humans, mice, pigs and chickens in which the highest expression of Lpin2 was found in liver, the highest expression of Lpin2 was found in perirenal and tail fats in GLT and in tail fat in STH. Based on the research reported, it would be likely that lipin2 plays crucial roles either by PAP activity or by regulator in TAG biosynthesis in sheep, and the low expression of Lpin3 in adipose tissues particularly in STH may mean its low PAP activity in TAG biosynthesis.
Studies suggested that Lpin2 mRNA is liver-enriched and has the significant PAP activity in liver, knockdown of Lpin2 markedly reduces the hepatocyte PAP activity and suppresses TAG synthesis (Gropler et al., 2009). Fld mice which lack lipin1 still have significant hepatic PAP activity (Donkor et al., 2007), and hepatocytes isolated from adult fld mice were shown to secrete VLDL at increased rates compared with wild-type hepatocytes (Burgdorf et al., 2009; Khalil et al., 2009), the results above could be attributable to compensatory upregulation of Lpin3 mRNA (Donkor et al., 2007;2009) and substantial expression of hepatic lipin2 protein (Gropler et al., 2009), which also suggest that lipin2 and/or lipin3 are capable of promoting hepatocyte VLDL synthesis and secretion. In our study, Lpin2 expression in liver was much lower than those in adipose tissues in GLT, although Lpin2 mRNA level in liver was similar to that in tail fat in STH. To some extent, the differences may be related with species differences. It is undetermined whether strong fat deposits in GLT influences or inhibits Lpin2 expression in liver. Meanwhile, the regulation mechanism of lipin2 in liver is complex, the compensatory increasing of lipin2 protein is under the translational control, and is independent of the steady-state Lpin2 mRNA levels in fld hepatocytes (Gropler et al., 2009). Anyway, the relative high expression of Lpin2 in liver in STH, as well as the highest expression of Lpin3 in liver in both breeds of sheep may mean their important functions in hepatic TAG synthesis and VLDL secretion.
Consistent with the studies reported (Donkor et al., 2007), Lpin3 rather than Lpin2 was highly expressed in small intestine in sheep. Some studies demonstrated that lipin1 can maintain nuclear/ER membrane morphology in mammalian or lower organisms (Golden et al., 2009; Gorjánácz and Mattaj, 2009; Sasser et al., 2012). Meanwhile, the role of lipin3 in small intestine seems not to be related with the TAG biosynthesis (Harris and Finck, 2011). In view of the complex physiological interactions among lip in proteins (Donkor et al., 2009; Gropler et al., 2009; Dwyer et al., 2012), raised the possibility that lipin3 functions in a separate or cooperative way with lipin1 in maintaining the intestinal membrane morphology in sheep, however, need to be proved.
Human Lpin2 mRNA is expressed in placenta (Donkor et al., 2007), chicken Lpin2 mRNA is ovary enriched (Zhang et al., 2014), which mean that lipin2 might involve in reproduction biology. In our study, Lpin2 and Lpin3 mRNA both were found in reproductive organs (testis and ovary), and Lpin2 expression in reproductive organs (testis and ovary) in GLT was higher than that in STH, and those were not significantly different from Lpin3 expression between two breeds. Lpin2 and Lpin3 expression both showed no significant differences between males and females as well between testis and ovary. The roles of lipin2 and lipin3 in reproduction biology in sheep are unclear. Unlike lipin1 which with high expression in skeletal muscle and functions in energy metabolism (Phan and Reue, 2005), the low abundance of Lpin2 and Lpin3 in femoral biceps may suggest that they may not be involved in energy metabolism in sheep.
Lpin2 expression levels reached a peak at 10 months of age in two breeds of sheep, while the lowest expression occurred at 2 months of age in STH and at 8 months of age in GLT. Studies suggested that the Lpin2 mRNA expression in liver is upregulated in response to fasting, diet-induced obesity in mice (Donkor et al., 2009; Gropler et al., 2009), which means Lpin2 mRNA expression can be induced in some context. It is noteworthy that 10 months of age was just in September, the mild climate and abundant forage in this month in Northern China benefit fat deposits, and possibly induced Lpin2 mRNA expression. Meanwhile, 2 and 8 months of age were in January and July, the cold climate in January and the hot weather in July may impact sheep feed intake and further inhibit fatty synthesis. Notably, the high expression was also found at 4 months of age in GLT, this stage was in March and it was cold and forage was scarce. Studies suggested that lipin2 can be a regulator in preadipocytes (Grimsey et al., 2008), consequently, the high expression of Lpin2 at 4 months of age in GLT may indicate its regulation role in preadipocyte formation.
In this study, we found the highest expression of Lpin3 occurred at 4 months of age in STH and at 10 months of age in GLT, the lowest levels occurred at 12 months of age in STH and at 8 months of age in GLT. In addition, Lpin3 mRNA was also highly expressed at 2 to 4 months of age in GLT, and Lpin3 mRNA levels in liver, kidney and small intestine were high at 2 to 4 months of age accordingly, which appear to suggest that lipin3 plays a function in these tissues mainly in early growth period. Compared with GLT, the expression levels of Lpin3 in STH were low at 10 to 12 months of age. Studies suggested that the PAP specific activity of lipin3 are far lower than that of lipin1 (Donkor et al., 2007), therefore, the low expression of Lpin3 at 10 to 12 months of age may indicate its low PAP activity. However, because Lpin3 expression levels in perirenal and tail fats in GLT were higher than those in STH, combined with the high expression of Lpin3 at 10 to 12 months of age in GLT, it’s likely that lipin3 plays PAP activity mainly in active lipid metabolism such as GLT.
In this study, we found that Lpin2 expression in tissues showed mostly negative correlations with traits in GLT, however, which showed prominently positive correlations in STH. In spite of the high expression of Lpin2 in perirenal and tail fats particularly at 4 and 10 months of age in GLT, but showed no significant correlations with slaughter and tail traits, which could be explained by the high expression in perirenal and tail fats yet low traits values at 4 months of age in GLT, however, the high expression of Lpin2 in adipose tissues and the high traits values at 10 months of age may indicate lipin2 promotes fat deposits by PAP activity. The expression of Lpin2 in liver was lower than those in adipose tissues, which may be the reason for the significant negative correlations between low expression in liver and strong fat deposits in GLT. Moreover, the significant positive correlations between the expression of Lpin2 in reproductive organs (esp. testis) and traits may indicate its reproductive biology function and furthermore promotes fat deposits and weight gain. The lowest expression of Lpin2 in femoral biceps caused the significant negative correlations with traits in GLT. By contrast, the average expression of Lpin2 was high at 8 to 12 months of age, which was synchronous with the vigorous growth stages of 8 to 12 months of age in STH, therefore the high Lpin2 expression in liver, kidney, perirenal and tail fats, as well as in femoral biceps exhibited the significant positive correlations with the traits, and ultimately promoted the weight gain in STH. Unlike GLT, the expression of Lpin2 in reproductive organs (testis and ovary) showed no significant correlations with traits in STH, which may due to the breed differences. The non-significant correlations between Lpin2 expression in small intestine and traits in both breeds may mean lipin2 does not function in small intestine in sheep.
As is shown in Table 5, Lpin3 expression in tissues showed mostly positive correlations with traits in GLT. Lpin3 was expressed at high levels highly in liver, small intestine, perirenal and tail fats in GLT, the significant positive correlations between Lpin3 expression in liver and absolute tail fat weight, small intestine and dressing percentage, perirenal fat and tail width, body weight, carcass weight as well as absolute tail fat weight may indicate lipin3 promotes the fat deposits and weight gain by PAP activity or transcriptional co-activators in GLT. Compared with GLT, Lpin3 expression in tissues showed mostly negative correlations in STH, Unlike GLT in which the high expression of Lpin3 occurred at 10 to 12 months of age, the highest expression of Lpin3 occurred at 4 months of age and with low expression at 6 to 12 months of age in STH, relative low Lpin3 expression as well as relative high phenotype values at 8 to 12 months of age could explain the significant negative correlations between expression and traits in STH. Moreover, Lpin3 expression was low in fat tissues, which may indicate no significant correlations between Lpin3 expression in adipose tissues and traits in STH.
ACKNOWLEDGMENTS
The authors would like to thank the staff of the Laboratory of Animal Genetics, Breeding and Reproduction, Shanxi Agricultural University. The work was supported by the projects of the Chinese National Natural Science Foundation (31372292).
Figure 1
Effect of breed by age on the abundance of ovine Lpin2 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed.
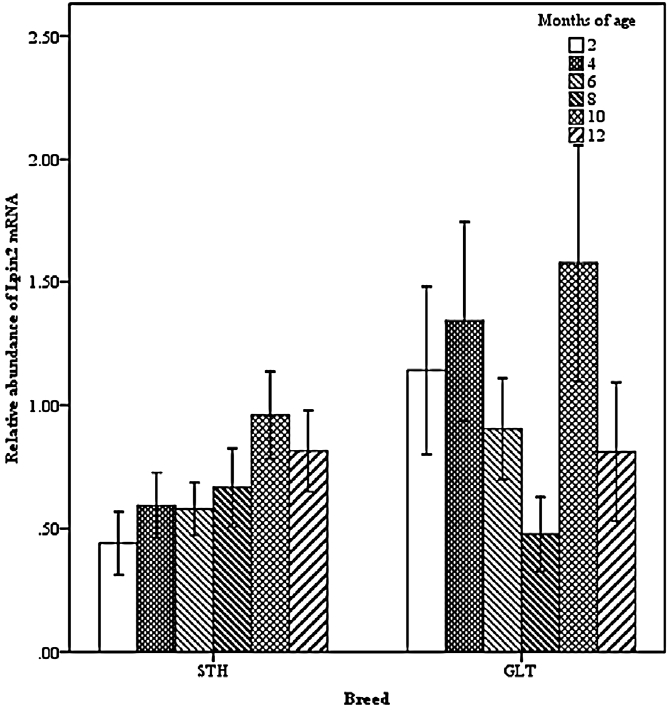
Figure 2
Effect of breed by tissue on the abundance of ovine Lpin2 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed; LIVE, liver; KIDN, kidney; SMIN, small intestine; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat.
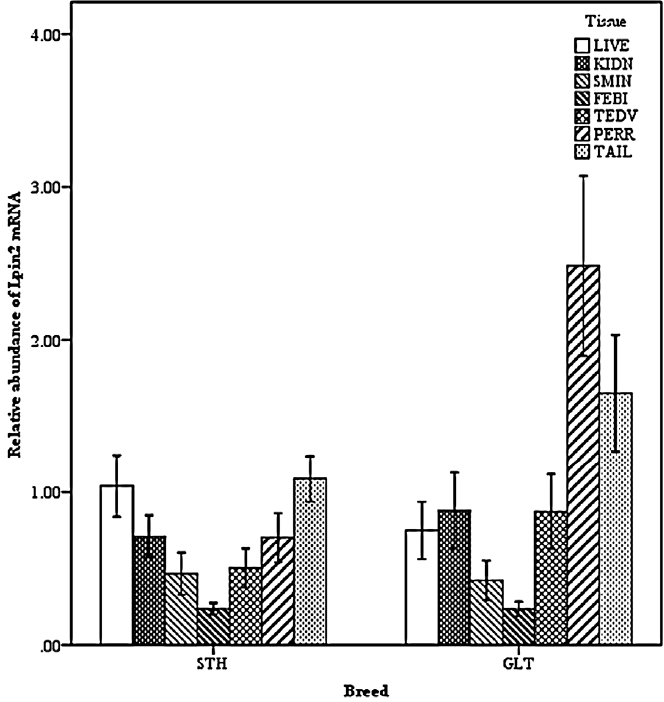
Figure 3
Effect of tissue by age on the abundance of ovine Lpin2 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed; LIVE, liver; KIDN, kidney; SMIN, small intestine; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat.

Figure 4
Effect of breed by age on the abundance of ovine Lpin3 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed.
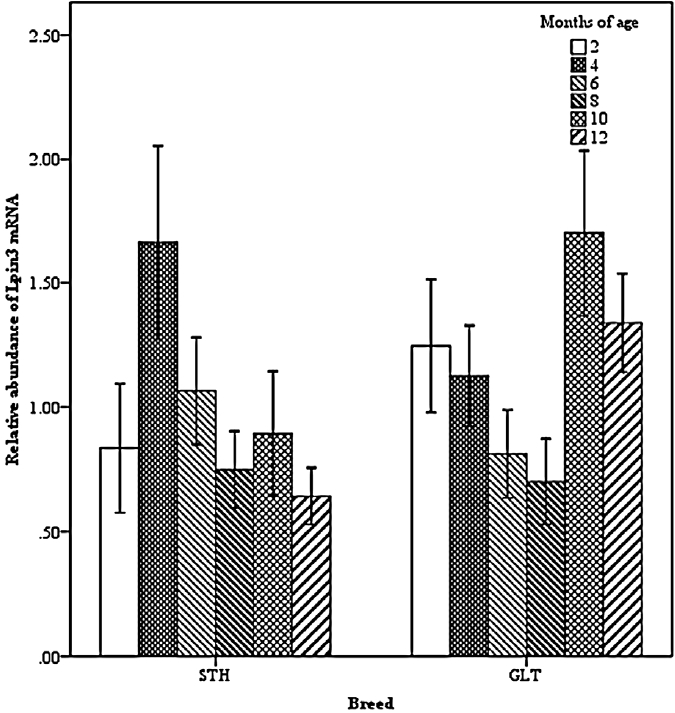
Figure 5
Effect of breed by tissue on the abundance of ovine Lpin3 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed; LIVE, liver; KIDN, kidney; SMIN, small intestine; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat.
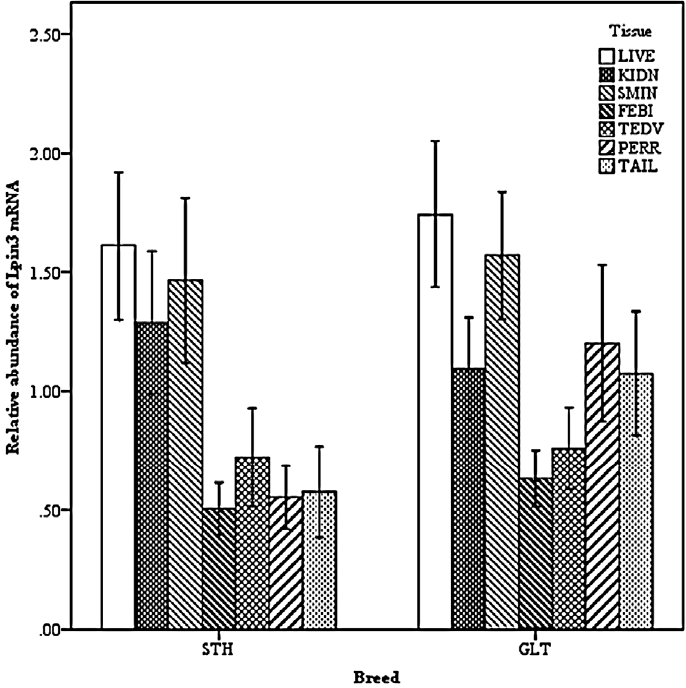
Figure 7
Effect of tissue by age on the abundance of ovine Lpin3 mRNA. STH, Small Tailed Han; GLT, Guangling Large Tailed; LIVE, liver; KIDN, kidney; SMIN, small intestine; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat.
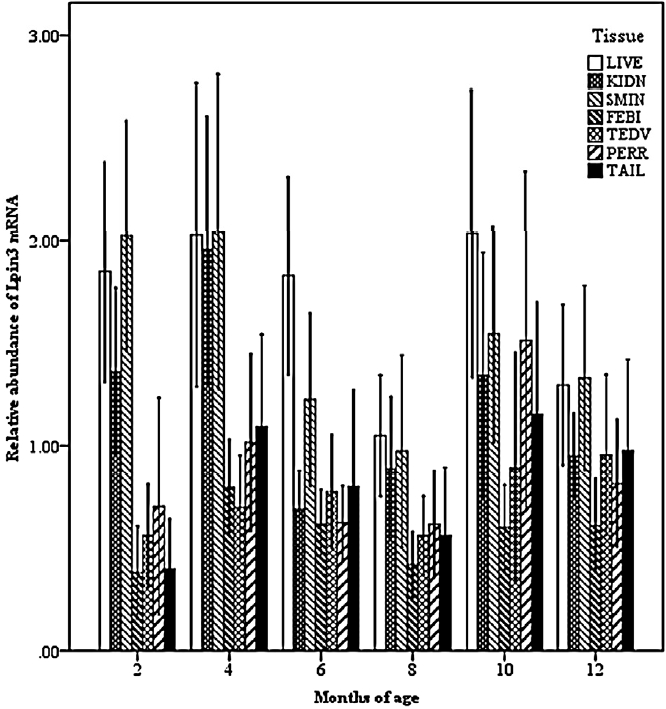
Table 1
Primer sequence, annealing temperature and PCR product size
Table 2
Analysis of variance for the mRNA abundance of ovine Lpin2 and Lpin3
| Source of variation | Sum of squares | Degree of freedom | F values | Significance | |||
|---|---|---|---|---|---|---|---|
|
|
|
|
|||||
| Lpin2 | Lpin3 | Lpin2 | Lpin | Lpin2 | Lpin3 | ||
| Corrected model | 317.529a | 239.687a | 66 | 8.114 | 6.424 | *** | *** |
| Intercept | 483.039 | 748.304 | 1 | 814.629 | 1,323.743 | *** | *** |
| Breed | 22.549 | 5.2 | 1 | 38.028 | 9.199 | *** | ** |
| Sex | 0.162 | 0.626 | 1 | 0.274 | 1.107 | NS | NS |
| MOA | 28.702 | 32.412 | 5 | 9.681 | 11.467 | *** | *** |
| Tissue | 131.954 | 99.495 | 6 | 37.089 | 29.334 | *** | *** |
| Breed×sex | 0.607 | 0.586 | 1 | 1.024 | 1.036 | NS | NS |
| Breed×MOA | 21.95 | 42.344 | 5 | 7.403 | 14.981 | *** | *** |
| Breed×tissue | 66.493 | 11.865 | 6 | 18.69 | 3.498 | *** | ** |
| Sex×MOA | 6.306 | 10.315 | 5 | 2.127 | 3.649 | NS | ** |
| Sex×tissue | 4.493 | 7.034 | 6 | 1.263 | 2.074 | NS | NS |
| Tissue×MOA | 29.68 | 33.219 | 30 | 1.668 | 1.959 | * | ** |
| Error | 354.588 | 338.046 | 598 | ||||
| Total | 1,166.079 | 1,322.672 | 665 | ||||
| Corrected total | 672.117 | 577.733 | 664 | ||||
Table 3
Abundance of ovine Lpin2 and Lpin3 mRNA
Table 4
Associations between Lpin2 mRNA expression and traits in GLT and STH
| Traits | Breed | Abundance of Lpin2 mRNA | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
|
|
||||||||||
| LIVE | KIDN | SMIN | FEBI | TEOV | PERR | TAIL | Testis | Ovary | ||
| TL | GLT | −0.342* | −0.002 | −0.202 | −0.214 | 0.355* | −0.135 | −0.289 | 0.645** | −0.139 |
| STH | 0.429** | 0.402** | 0.018 | 0.514** | −0.018 | 0.359* | 0.354* | −0.076 | 0.079 | |
| TW | GLT | −0.446** | −0.037 | −0.11 | −0.157 | 0.304* | −0.139 | −0.058 | 0.639** | −0.216 |
| STH | 0.505** | 0.305* | 0.041 | 0.455** | −0.181 | 0.540** | 0.448** | −0.194 | −0.03 | |
| BW | GLT | −0.441** | 0.004 | −0.085 | −0.364* | 0.386** | 0.085 | −0.113 | 0.7** | −0.168 |
| STH | 0.511** | 0.306* | −0.126 | 0.568** | 0.071 | 0.366* | 0.612** | 0.145 | −0.204 | |
| CW | GLT | −0.439** | −0.026 | −0.1 | −0.381* | 0.386** | 0.024 | −0.117 | 0.722** | −0.073 |
| STH | 0.538** | 0.279 | −0.099 | 0.533** | −0.017 | 0.412** | 0.589** | 0.029 | −0.237 | |
| DP | GLT | −0.167 | −0.073 | −0.127 | −0.227 | 0.188 | −0.025 | −0.063 | 0.412 | 0.166 |
| STH | 0.299 | 0.003 | 0.055 | 0.144 | −0.179 | 0.335* | 0.237 | −0.186 | −0.21 | |
| ATW | GLT | −0.447** | 0.149 | −0.08 | −0.271 | 0.460** | 0.056 | −0.072 | 0.774** | −0.121 |
| STH | - | - | - | - | - | - | - | - | - | |
| RTW | GLT | −0.460** | 0.141 | −0.1 | −0.146 | 0.441** | −0.004 | −0.06 | 0.73** | −0.013 |
| STH | - | - | - | - | - | - | - | - | - | |
GLT, Guangling Large Tailed; STH, Small Tailed Han; LIVE, liver; KIDN, kidney; SMIN, small intestines; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat; TL, tail length; TW, tail width; BW, body weight; CW, carcass weight; DP, dressing percentage; ATW, absolute tail fat weight; RTW, relative tail fat weight.
Table 5
Associations between Lpin3 mRNA expression and traits in GLT and STH
| Traits | Breed | Abundance of Lpin3 mRNA | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
|
|
||||||||||
| LIVE | KIDN | SMIN | FEBI | TEOV | PERR | TAIL | Testis | Ovary | ||
| TL | GLT | 0.136 | −0.188 | −0.007 | −0.034 | −0.045 | 0.244 | 0.09 | −0.173 | 0.097 |
| STH | −0.349* | −0.022 | −0.274 | −0.261 | 0.084 | 0.031 | −0.017 | 0.023 | 0.125 | |
| TW | GLT | 0.212 | 0.02 | 0.173 | 0.129 | 0.121 | 0.317* | 0.372* | −0.015 | 0.245 |
| STH | −0.322* | −0.083 | −0.281 | −0.205 | 0 | 0.139 | 0.264 | −0.042 | 0.041 | |
| BW | GLT | 0.24 | −0.095 | 0.08 | 0.249 | 0.181 | 0.408** | 0.235 | −0.017 | 0.408 |
| STH | −0.26 | −0.126 | −0.284 | −0.264 | 0.025 | 0.156 | 0.091 | −0.017 | 0.105 | |
| CW | GLT | 0.29 | −0.072 | 0.193 | 0.21 | 0.236 | 0.379* | 0.231 | 0.013 | 0.413 |
| STH | −0.28 | −0.146 | −0.346* | −0.294 | 0.02 | 0.1 | 0.078 | −0.033 | 0.108 | |
| DP | GLT | 0.286 | 0.018 | 0.361* | −0.029 | 0.186 | 0.065 | 0.076 | 0.099 | 0.201 |
| STH | −0.304* | −0.244 | −0.460** | −0.292 | 0.034 | −0.116 | 0.018 | −0.044 | 0.111 | |
| ATW | GLT | 0.296* | −0.037 | 0.095 | 0.173 | 0.035 | 0.411** | 0.275 | −0.097 | 0.198 |
| STH | - | - | - | - | - | - | - | - | - | |
| RTW | GLT | 0.209 | −0.033 | −0.02 | 0.119 | −0.122 | 0.29 | 0.244 | −0.23 | −0.011 |
| STH | - | - | - | - | - | - | - | - | - | |
GLT, Guangling Large Tailed; STH, Small Tailed Han; LIVE, liver; KIDN, kidney; SMIN, small intestines; FEBI, femoral biceps; TEOV, testis and ovary; PERR, perirenal fat; TAIL, tail fat; TL, tail length; TW, tail width; BW, body weight; CW, carcass weight; DP, dressing percentage; ATW, absolute tail fat weight; RTW, relative tail fat weight.
REFERENCES
Burgdorf C, Hänsel L, Heidbreder M, Jöhren O, Schütte F, Schunkert H, Kurz T. 2009. Suppression of cardiac phosphatidatephosphohydrolase1 activity and lipin mRNA expression in Zucker diabetic fatty rats and humans with type 2 diabetes mellitus. Biochem Biophys Res Commun 390:165–170.


Chen Z, Gropler MC, Mitra MS, Finck BN. 2012. Complex interplay between the lipin1 and the hepatocyte nuclear factor 4a (HNF4a) pathways to regulate liver lipid metabolism. PloS One 7:e51320



Donkor J, Sariahmetoglu M, Dewald J, Brindley DN, Reue K. 2007. Three mammalian lipins act as phosphatidate phosphatases with distinct tissue expression patterns. J Biol Chem 282:3450–3457.


Donkor J, Zhang P, Wong S, O’Loughlin L, Dewald J, Kok BP, Brindley DN, Reue K. 2009. A conserved serine residue is required for the phosphatidate phosphatase activity but not the transcriptional coactivator functions of lipin-1 and lipin-2. J Biol Chem 284:29968–29978.



Dwyer JR, Donkor J, Zhang P, Csaki LS, Vergnes L, Lee JM, Dewald J, Brindley DN, Atti E, Tetradis S, Yoshinaga Y, De Jong PJ, Fong LG, Young SG, Reue K. 2012. Mouse lipin-1 and lipin-2 cooperate to maintain glycerolipid homeostasis in liver and aging cerebellum. Proc Natl Acad Sci USA 109:E2486–2495.



Golden A, Liu J, Cohen-Fix O. 2009. Inactivation of the C. Elegans lipin homolog leads to ER disorganization and to defects in the breakdown and reassembly of the nuclear envelope. J Cell Sci 122:1970–1978.



González CR, Novelle MG, Caminos JE, Vázquez MJ, Luque RM, López M, Nogueiras R, Diéguez C. 2012. Regulation of lipin1 by nutritional status, adiponectin, sex and pituitary function in rat white adipose tissue. Physiol Behav 105:777–783.


Gorjánácz M, Mattaj IW. 2009. Lipin is required for efficient breakdown of the nuclear envelope in Caenorhabditis elegans. J Cell Sci 122:1963–1969.


Grimsey N, Han GS, O’Hara L, Rochford JJ, Carman GM, Siniossoglou S. 2008. Temporal and spatial regulation of the phosphatidate phosphatases lipin 1 and 2. J Biol Chem 283:29166–29174.



Gropler MC, Harris TE, Hall AM, Wolins NE, Gross RW, Han X, Chen Z, Finck BN. 2009. Lipin 2 is a liver-enriched phosphatidate phosphohydrolase enzyme that is dynamically regulated by fasting and obesity in mice. J Biol Chem 284:6763–6772.



Han GS, Wu WI, Carman GM. 2006. The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme. J Biol Chem 281:9210–9218.



Harris TE, Finck BN. 2011. Dual function lipin proteins and glycerolipid metabolism. Trends Endocrinol Metab 22:226–233.



He X, Xu X, Liu B. 2009. Molecular characterization, chromosomal localization and association analysis with backfat thickness of porcine LPIN2 and LPIN3. Mol Biol Rep 36:1819–1824.


Herlin T, Fiirgaard B, Bjerre M, Kerndrup G, Hasle H, Bing X, Ferguson PJ. 2013. Efficacy of anti-IL-1 treatment in Majeed syndrome. Ann Rheum Dis 72:410–413.



Huang H, Gao Q, Peng X, Choi SY, Sarma K, Ren H, Morris AJ, Frohman MA. 2011. piRNA-associated germline nuage formation and spermatogenesis require MitoPLD profusogenic mitochondrial-surface lipid signaling. Dev Cell 20:376–387.



Khalil BM, Sundaram M, Zhang HY, Links PH, Raven JF, Manmontri B, Sariahmetoglu M, Tran K, Reue K, Brindley DN, Yao Z. 2009. The level and compartmentalization of phosphatidate phosphatase-1 (lipin-1) control the assembly and secretion of hepatic VLDL. J Lipid Res 50:47–58.


Livak KJ, Schmittgen TD. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta DeltaC(T)). Methods 25:402–408.


Michot C, Hubert L, Brivet M, De Meirleir L, Valayannopoulos V, Müller-Felber W, Venkateswaran R, Ogier H, Desguerre I, Altuzarra C, Thompson E, Smitka M, Huebner A, Husson M, Horvath R, Chinnery P, Vaz FM, Munnich A, Elpeleg O, Delahodde A, deKeyzer Y, deLonlay P. 2010. LPIN1 gene mutations: a major cause of severe rhabdomyolysis in early childhood. Hum Mutat 31:E1564–1573.


Mitra MS, Chen Z, Ren H, Harris TE, Chambers KT, Hall AM, Nadra K, Klein S, Chrast R, Su X, Morris AJ, Finck BN. 2013. Mice with an adipocyte-specific lipin 1 separation-of-function allele reveal unexpected roles for phosphatidic acid in metabolic regulation. Proc Natl Acad Sci USA 110:642–647.



Péterfy M, Harris TE, Fujita N, Reue K. 2010. Insulin-stimulated interaction with 14-3-3 promotes cytoplasmic localization of lipin-1 in adipocytes. J Biol Chem 285:3857–3864.



Péterfy M, Phan J, Reue K. 2005. Alternatively spliced lipin isoforms exhibit distinct expression pattern, subcellular localization, and role in adipogenesis. J Biol Chem 280:32883–32889.


Péterfy M, Phan J, Xu P, Reue K. 2001. Lipodystrophy in the fld mouse results from mutation of a new gene encoding a nuclear protein, lipin. Nat Genet 27:121–124.


Phan J, Peterfy M, Reue K. 2004. Lipin expression preceding peroxisome proliferator-activated receptor-gamma is critical for adipogenesis in vivo and in vitro. J Biol Chem 279:29558–29564.


Reue K, Xu P, Wang XP, Slavin BG. 2000. Adipose tissue deficiency, glucose intolerance, and increased atherosclerosis result from mutation in the mouse fatty liver dystrophy (fld) gene. J Lipid Res 41:1067–1076.


Sanderson LM, Degenhardt T, Koppen A, Kalkhoven E, Desvergne B, Müller M, Kersten S. 2009. Peroxisome proliferator-activated receptor beta/delta (PPARbeta/delta) but not PPARalpha serves as a plasma free fatty acid sensor in liver. Mol Cell Biol 29:6257–6267.



Sasser T, Qiu QS, Karunakaran S, Padolina M, Reyes A, Flood B, Smith S, Gonzales C, Fratti RA. 2012. Yeast lipin 1 orthologue pah1p regulates vacuole homeostasis and membrane fusion. J Biol Chem 287:2221–2236.



Scott RA, Lagou V, Welch RP, Wheeler E. 2012. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat Genet 44:991–1005.








 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





