 |
 |
| Anim Biosci > Volume 36(10); 2023 > Article |
|
Abstract
Objective
╬▒S1-Casein is more closely associated with milk allergic reaction than other milk protein components. microRNA (miRNA) is a class of small non-coding RNAs that modulate multiple biological progresses by the target gene. However, the post-transcriptional regulation of ╬▒S1-casein expression by miRNA in ruminants remains unclear. This study aims to explore the regulatory roles of miR-380-3p on ╬▒S1-casein synthesis in goat mammary epithelial cells (GMEC).
Methods
╬▒S1-Casein gene and miR-380-3p expression was measured in dairy goat mammary gland by quantitative real-time polymerase chain reaction (qRT-PCR). miR-380-3p overexpression and knockdown were performed by miR-380-3p mimic or inhibitor in GMEC. The effect of miR-380-3p on ╬▒S1-casein synthesis was detected by qRT-PCR, western blot, luciferase and chromatin immunoprecipitation assays in GMEC.
Results
Compared with middle-lactation period, ╬▒S1-casein gene expression is increased, while miR-380-3p expression is decreased during peak-lactation of dairy goats. miR-380-3p reduces ╬▒S1-casein abundance by targeting the 3ŌĆ▓-untranslated region (3ŌĆ▓UTR) of ╬▒S1-casein mRNA in GMEC. miR-380-3p enhances ╬▓-casein expression and signal transducer and activator of transcription 5a (STAT5a) activity. Moreover, miR-380-3p promotes ╬▓-casein abundance through target gene ╬▒S1-casein, and activates ╬▓-casein transcription by enhancing the binding of STAT5 to ╬▓-casein gene promoter region.
Milk protein is a dietary protein source that supplies essential amino acids and bioactive peptides for humans, mainly composed of caseins and whey proteins. Caseins account for about 80% of the total milk protein, and are divided into ╬▒S1-, ╬▒S2-, ╬▓- and ╬║-casein. Despite its benefits, milk protein also carries a risk of food allergies for many people, especially for infants and children [1]. The global prevalence of milk protein allergy was reported to be as high as 2% to 7.5% [2]. Among the numerous components of milk protein, ╬▒S1-casein is most prone to cause allergic reactions, and approximately 65% of patients with milk allergies are caused by ╬▒S1-casein [3]. Goat milk contains a variable content of ╬▒S1-casein (4% to 26% of milk protein) compared to cow milk (36% to 40%) [4]. Therefore, the method of regulating ╬▒S1-casein synthesis to inhibit the allergic potential of milk protein is necessary to be elucidated.
MicroRNA (miRNA) is a class of small non-coding RNA of about 22 nucleotides in length, which regulate mRNA expression by targeting the 3ŌĆ▓-untranslated region (3ŌĆ▓UTR) of gene transcripts. The pairing between miRNA seed site sequence (2nd to 8th nucleotides at the 5ŌĆ▓ terminal) and target genes is considered a key feature for mammalian miRNA to recognize its targets. The abundance of miR-380-3p is involved in the prevention and treatment of a variety of human diseases. miR-380-3p participates in intracorporal neurodegeneration induced by neurotoxicants via specificity protein 3 signaling pathway [5]. High expression of miR-380-3p inhibits cell proliferation and invasion by targeting oncogene Kruppel-like factor 4 in triple negative breast cancer cells [6]. miR-380-3p increases the anti-proliferative and pro-apoptotic effects of doxorubicin in multiple models of high-risk neuroblastoma [7].
To address the milk allergy caused by ╬▒S1-casein in ruminants, it is urgent to investigate the regulatory mechanism of miRNAs on ╬▒S1-casein synthesis. Although multiple studies have revealed that miR-380-3p plays the pivotal role in a variety of biological processes, whether it could modulate ruminant milk protein expression is unknown. ╬▒S1-Casein gene was targeted by miR-380-3p in dairy goats or cows via TargetScan database ( http://www.targetscan.org/) prediction. Hence, the aim of our study was to explore the molecular mechanism by which miR-380-3p controls milk protein synthesis through the target gene in goat mammary epithelial cells (GMEC). In this study, we revealed that miR-380-3p reduced ╬▒S1-casein synthesis and enhanced ╬▓-casein abundance in GMEC, which supplies a novel approach to weaken milk allergy potential and promote milk nutritional quality in ruminants.
The animal study protocol was approved by the Institutional Animal Care and Use Committee, Northwest A&F University, YangLing, China (protocol code 15ŌĆō516).
Mammary gland samples were obtained from 3-year-old healthy Xinong Saanen dairy goats (n = 5) during early, peak, middle and end lactation periods (15, 60, 120, and 270 days postpartum, respectively). GMEC were isolated from three dairy goats at peak lactation period. Cells were cultured in the incubator at 37┬░C and 5% CO2. The culture medium contained 90% DMEM/F12 (SH30023; Hyclone, Logan, UT, USA) medium and 10% fetal bovine serum (10099141; Gibco, Gaithersburg, MD, USA), with 5 ╬╝g/mL bovine insulin (16634; Sigma, St. Louis, MO, USA), 1 ╬╝g/mL hydrocortisone (H0888; Sigma, USA), 10 ng/mL epidermal growth factor (PHG0311; Invitrogen, Carlsbad, CA, USA), 100 U/mL penicillin/streptomycin (080092569; Harbin Pharmaceutical Group, Harbin, China) added.
Primers for sequence cloning of goat ╬▒S1-casein gene (GenBank no. XM_018049127.1) and ╬▓-casein promoter region (GenBank no. NC_030813.1) are shown in Supplementary Table S1. The template of coding sequence and 3ŌĆ▓UTR region cloning of ╬▒S1-casein was cDNA obtained from goat mammary gland. The template of ╬▓-casein promoter region cloning was goat blood genomic DNA. pcDNA3.1-╬▒S1-casein vector was constructed using goat ╬▒S1-casein coding sequence and pcDNA3.1 plasmid. psi-WT-╬▒S1-Casein vector was produced using ╬▒S1-casein 3ŌĆ▓UTR sequence and psiCHECK2 plasmid. ╬▒S1-Casein 3ŌĆ▓UTR site-directed mutant vector psi-MUT-╬▒S1-Casein was produced using overlapping polymerase chain reaction (PCR) method. pGL3-WT-╬▓-Casein vector was constructed using ╬▓-casein promoter sequence and pGL3-Basic plasmid. Similarly, ╬▓-casein promoter site-directed mutant vector was performed and named pGL3-MUT-╬▓-Casein. All constructed vectors were sequenced to confirm the sequences.
╬▒S1-Casein 3ŌĆ▓UTR was targeted by miR-380-3p using TargetScan database prediction., The experimental treatments were carried out when cell confluence reached 80%. For miR-380-3p overexpression, cells were transfected with miR-380-3p mimic or the mimic negative control (NC) (50 nM; RiboBio, Guangzhou, China) using Lipofectamine RNAiMAX (13778150; Invitrogen, USA). Similarly, for miR-380-3p knockdown, GMEC were treated with miR-380-3p inhibitor or the negative control inhibitor NC (100 nM; RiboBio, China). We silenced ╬▒S1-casein gene by specific small interference RNA (si-╬▒S1-Casein; GenePharma, Shanghai, China). The sequences of si-╬▒S1-Casein and the negative control si-NC are shown in Supplementary Table S2. GMEC were co-treated with si-╬▒S1-Casein (100 nM) and miR-380-3p inhibitor (100 nM) using Lipofectamine RNAiMAX. ╬▒S1-Casein overexpression was carried out by pcDNA3.1-╬▒S1-casein plasmid. Cells were co-treated with pcDNA3.1-╬▒S1-casein and miR-380-3p mimic (50 nM) using X-treme GENE HP Regent (06366236001; Roche, Mannheim, Germany). GMEC were collected after 48 h treatment to extract RNA and protein.
Total RNA of cells was extracted by RNAiso Plus (9109; Takara, Shiga, Japan). The quality of total RNA was detected by NanoDrop2000 spectrophotometer (Thermo Scientific, Waltham, MA, USA). The optical density 260/280 of total RNA was 1.9 to 2.1, and the concentration was approximately 500 ng/╬╝L. For gene quantitative analysis, cDNA was synthesized by purified total RNA with PrimeScript RT kit (RR047A; Takara, Japan). Gene quantitative real-time PCR (qRT-PCR) was carried out with TB Green II kit (RR820A; Takara, Japan). The control genes were ubiquitously expressed transcript (UXT) and ribosomal protein S9 (RPS9). Primers for gene quantitative analysis are listed in Table 1. For miR-380-3p quantitative analysis, cDNA of mature miR-380-3p was produced by miRcute cDNA kit (KR211; Tiangen, Beijing, China). qRT-PCR of miR-380-3p were performed by miRcute q-PCR kit (FP411; Tiangen, China). The control for miR-380-3p was 5S rRNA. The quantitative primer sequence of miR-380-3p was 5ŌĆ▓-CGCTATGTAATGTGGTCC ACGTCT-3ŌĆ▓. The primer sequence of 5S rRNA was 5ŌĆ▓-TCTC AGAAGCTAAACAGGGTCG-3ŌĆ▓. All qRT-PCR experiments were carried out by Bio-Rad CFX96 analyzer (Hercules, CA, USA). Quantitative data were analyzed by 2ŌłÆ╬ö╬öCt method.
Total protein of cells was collected by radioimmunoprecipitation assay lysis regent (R0010; Solarbio, Beijing, China) with protease and phosphatase inhibitor (04693132001 and 04906845001; Roche, Germany). The concentration of total protein was detected by bicinchoninic acid assay kit (23227; Thermo Scientific, USA). The protein was separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and blotted onto the polyvinylidene fluoride membrane (Roche, Germany) with Bio-Rad Trans-Blot semidry transfer. Nonspecific sites were blocked using 5% skim milk (232100; BD Biosciences, Franklin Lakes, NJ, USA) at least 2 h, then incubated at least 12 h at 4┬░C with primary antibody: rabbit ╬▒S1-casein (SAB1401093; Sigma-Aldrich, USA; 1:1,000), rabbit ╬▓-casein (orb2053; Biorbyt, Cambridge, UK; 1:500), mouse signal transducer and activator of transcription 5a (STAT5a) (610191; BD Biosciences, USA; 1:1,000), mouse p-STAT5a (611964; BD Biosciences, USA; 1:500), or mouse ╬▓-actin (CW0096; CW Biotech, Beijing, China; 1:2,000). Secondary antibody was horseradish peroxidase-conjugated goat anti-mouse-immunoglobulin G (IgG) (CW0102; CW Biotech, China; 1:5,000) or goat anti-rabbit-IgG (CW0103; CW Biotech, China; 1:5,000). Blots were detected using chemiluminescent kit (1705061; Bio-Rad, USA). The blot intensity was analyzed by ImageJ ( http://imagej.nih.gov/ij/). Expression of caseins was normalized by comparison with ╬▓-actin, and expression of p-STAT5a was normalized to STAT5a.
For activity assay of 3ŌĆ▓UTR, GMEC were co-treated with ╬▒S1-casein 3ŌĆ▓UTR vectors and miR-380-3p mimic (50 nM) by X-treme GENE HP Regent. For activity assay of promoter, cells were co-treated with ╬▓-casein promoter vectors and miR-380-3p mimic (50 nM). ╬▓-Casein promoter vector was treated along with pRL-TK renilla vector as an internal control. Also, GMEC were treated with STAT5 inhibitor STAT5-IN-1 (50 ╬╝M, CAS 285986-31-4; MedChemExpress, NJ, USA) followed by miR-380-3p mimic (50 nM) and psi-WT-╬▓-Casein co-treatment. After 48 h treatment, cells were collected and lysed for luciferase assay. Data were measured using a Dual-Luciferase System (E1910; Promega, Madison, WI, USA) by Fluoroskan Ascent (Thermo Scientific, USA). The luciferase activity of 3ŌĆ▓UTR was analyzed as the ratio of renilla compared with firefly luciferase activity. The promoter activity was analyzed as the ratio of firefly compared with renilla activity.
The chromatin immunoprecipitation (ChIP) assay was carried out by Beyotime kit (P2048; Shanghai, China). Cells were cultured in 100-mm dish and treated with miR-380-3p mimic (50 nM) or STAT5-IN-1 (50 ╬╝M) for 48 h. Then, cells were crosslinked by 1% formaldehyde (final concentration) for 10 min at 37┬░C and harvested in cold PBS with protease and phosphatase inhibitor. Chromosomal DNA was sheared to lengths of 100 to 500 bp by Bioruptor Pico (Diagenode, Seraing, Belgium). The chromatin DNA-protein compound was incubated with protein A/G beads and rabbit p-STAT5 (9351S; Cell Signaling Technology, Danvers, MA, USA; 1:50) or control rabbit IgG (A7016; Beyotime, China; 1:50) for 12 h at 4┬░C. The primers containing STAT5 binding site on ╬▓-casein promoter region were shown in Table 1.
All experiments were repeated for at least for 3 biological replicates. Data were shown as mean┬▒standard error of the mean and analyzed by SPSS 20.0 software (IBM, Chicago, IL, USA). T-test was carried out with two group data, and one-way analysis of variance with Tukey test was carried out for multiple comparison. The difference was significant when p<0.05 (* p<0.05, ** p<0.01).
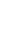
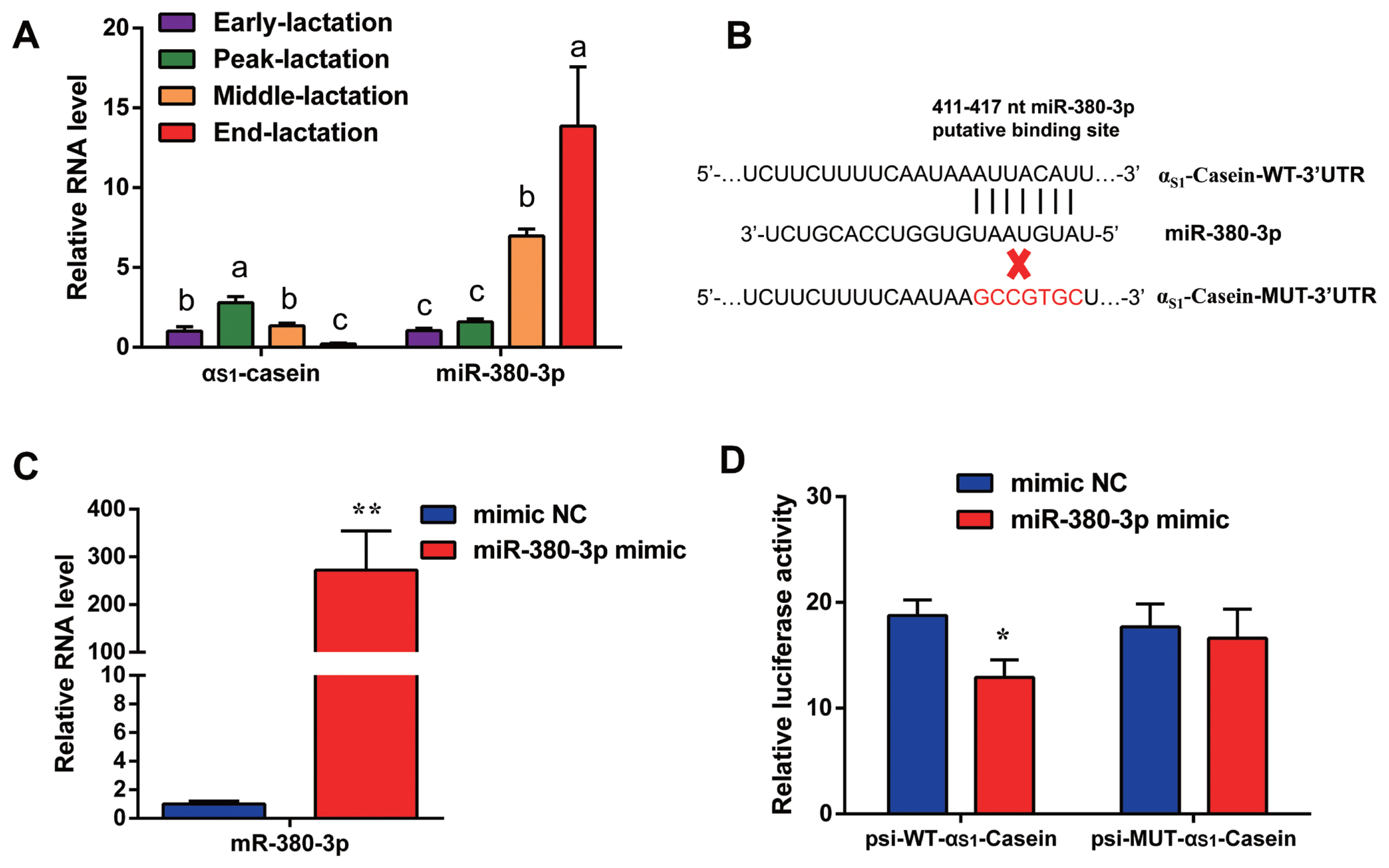
Quantitative result in goat mammary tissue showed that ╬▒S1-casein gene expression was highest in peak of lactation and lowest in end of lactation stage (Figure 1A). Based on TargetScan database prediction, miR-380-3p targets 3ŌĆ▓UTR of goat ╬▒S1-casein gene (Figure 1B). Expression of miR-380-3p was higher in end-lactation than other lactation stages. Specifically, ╬▒S1-casein gene expression was 2.13-fold higher in peak-lactation than middle-lactation stage (p<0.05), while miR-380-3p expression in middle-lactation was 4.39-fold higher than peak-lactation stage (p<0.05; Figure 1A).
To explore whether miR-380-3p did directly target 3ŌĆ▓UTR of ╬▒S1-casein, cells were treated with miR-380-3p mimic. Compared with mimic NC, miR-380-3p expression was up-regulated 272-fold in miR-380-3p mimic group (p<0.01; Figure 1C). The TargetScan database predicted that miR-380-3p binds to the bases at positions 411 to 417 of ╬▒S1-casein 3ŌĆ▓UTR (Figure 1B). Therefore, we detected dual-luciferase activity of wildtype and mutant plasmids after overexpressing miR-380-3p. Luciferase assay revealed that miR-380-3p could directly bind to specific site of ╬▒S1-casein 3ŌĆ▓UTR in dairy goats (Figure 1D).
Next, to confirm the effect of miR-380-3p on expression of ╬▒S1-casein, cells were transfected with mimic or inhibitor of miR-380-3p. Compared with mimic NC, expression of ╬▒S1-casein gene was significant reduced by an average of 64% in miR-380-3p mimic group (p<0.01; Figure 2A). Compared with inhibitor NC, expression of ╬▒S1-casein gene was effectively enhanced by an average of 78% in miR-380-3p inhibitor group (p<0.01; Figure 2B). Further, compared with control, miR-380-3p mimic could decrease ╬▒S1-casein expression by an average of 41% (p<0.05; Figure 2C). On the contrary, ╬▒S1-casein expression was increased by an average of 54% after miR-380-3p inhibition (p<0.05; Figure 2C). These finds suggest that miR-380-3p decreases ╬▒S1-casein expression which is strongly related to milk allergy potential.
To investigate whether miR-380-3p affects expression of other caseins, we measured the mRNA levels of ╬▒S2-casein, ╬▓-casein, and ╬║-casein gene. Results showed that overexpressing miR-380-3p increased ╬▓-casein gene expression (p<0.01; Figure 3A), but could not regulate ╬▒S2-casein and ╬║-casein mRNA levels. Similarly, knockdown of miR-380-3p decreased expression of ╬▓-casein gene as well (p<0.05; Figure 3B). On the other hand, ╬▒S2-casein and ╬║-casein mRNA levels had no significant difference between miR-380-3p inhibitor and inhibitor NC groups (Figure 3B). Then, we measured the expression of ╬▓-casein and STAT5a, a key transcription factor for milk protein synthesis. Results showed that miR-380-3p mimic markedly upregulated abundance of STAT5a phosphorylation (p-STAT5a) and ╬▓-casein (p<0.05; Figure 3C). Uniformly, miR-380-3p knockdown significantly reduced p-STAT5a and ╬▓-casein abundance (p<0.05; Figure 3C). Collectively, these results suggest that miR-380-3p decreases ╬▒S1-casein expression and upregulates ╬▓-casein expression, which is beneficial to hypoallergenic goat milk.
Our previous study discovered that ╬▒S1-casein gene decreased ╬▓-casein expression by inhibiting STAT5a activity in GMEC [8]. To confirm whether miR-380-3p increased ╬▓-casein expression through ╬▒S1-casein/STAT5a regulation axis, cells were co-transfected with miR-380-3p mimic and pcDNA3.1-╬▒S1-casein. Compared with mimic NC + pcDNA3.1-NC group, ╬▒S1-casein expression was markedly upregulated, while p-STAT5a and ╬▓-casein expression was decreased in mimic NC + pcDNA3.1-╬▒S1-casein group (p<0.05; Figure 4A, B). Compared with mimic NC + pcDNA3.1-╬▒S1-casein group, ╬▒S1-casein expression was significantly decreased, while p-STAT5a and ╬▓-casein expression was increased in miR-380-3p mimic + pcDNA3.1-╬▒S1-casein group (p<0.05; Figure 4A, B).
Similarly, silencing ╬▒S1-casein reduced ╬▒S1-casein abundance, with the increase of p-STAT5a and ╬▓-casein expression (p<0.05; Figure 4C, D). Furthermore, when we knocked down both ╬▒S1-casein and miR-380-3p, the increase in p-STAT5a and ╬▓-casein expression caused by ╬▒S1-casein silencing was abolished as a result of miR-380-3p inhibition (Figure 4C, D). Taken together, we deduce that miR-380-3p indirectly increases ╬▓-casein expression and STAT5 activity which leans upon ╬▒S1-casein reduction.
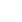
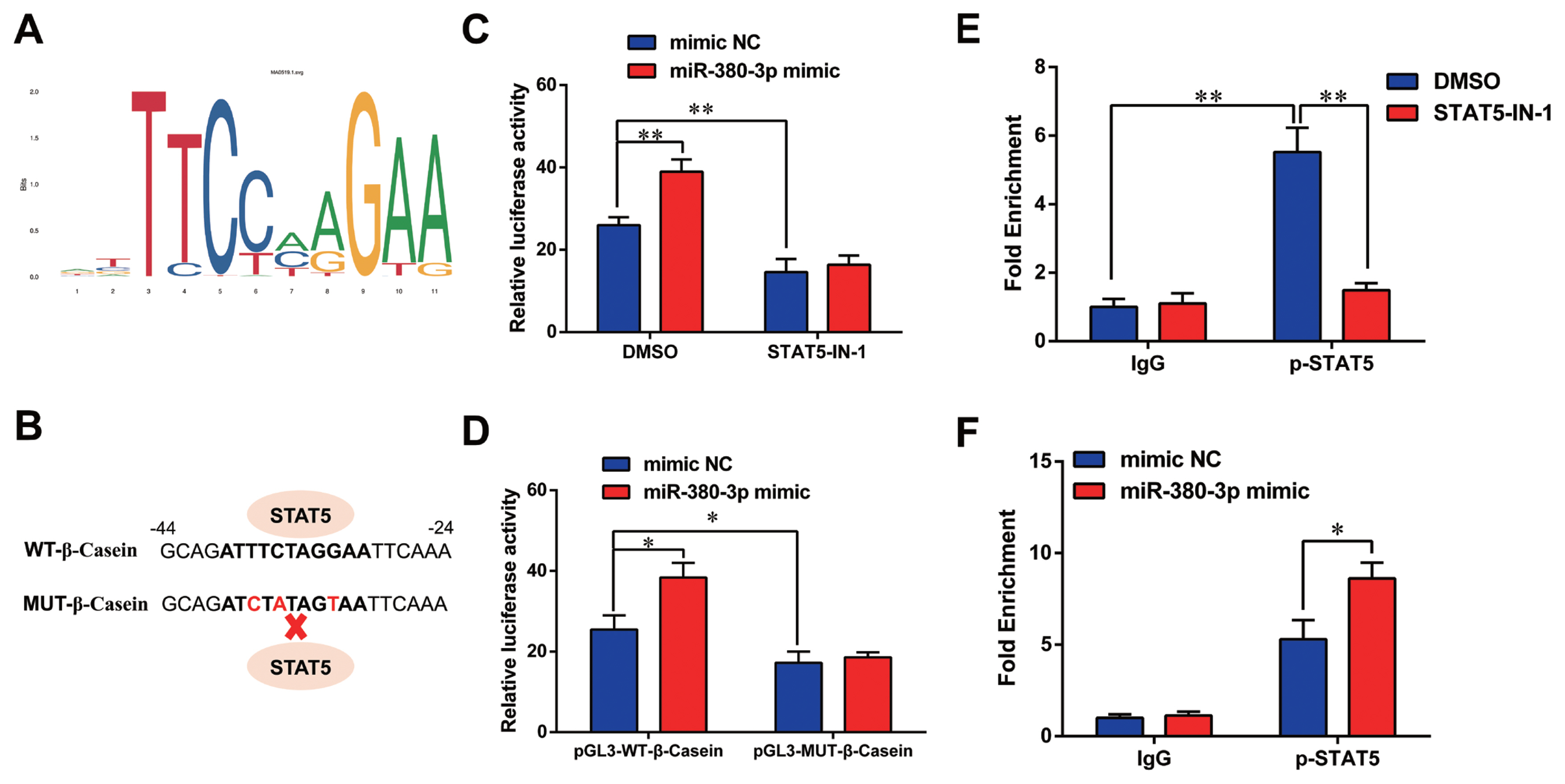
Previous studies have shown STAT5 enhances ╬▓-casein transcription by binding to ╬▓-casein promoter region in dairy goats [9]. Therefore, we determined whether miR-380-3p modulated ╬▓-casein transcription through STAT5. Figure 5A shows the conserved binding motif of transcription factor STAT5. We constructed the wild-type and mutant luciferase plasmids of ╬▓-casein promoter according to STAT5 binding site (Figure 5B). miR-380-3p obviously increased ╬▓-casein promoter activity, but STAT5 inhibitor reduced ╬▓-casein promoter activity regardless of miR-380-3p overexpression (p<0.01; Figure 5C). Consistently, compared with pGL3-WT-╬▓-Casein promoter group, pGL3-MUT-╬▓-Casein promoter activity was markedly weakened regardless of miR-380-3p overexpression (p<0.05; Figure 5D).
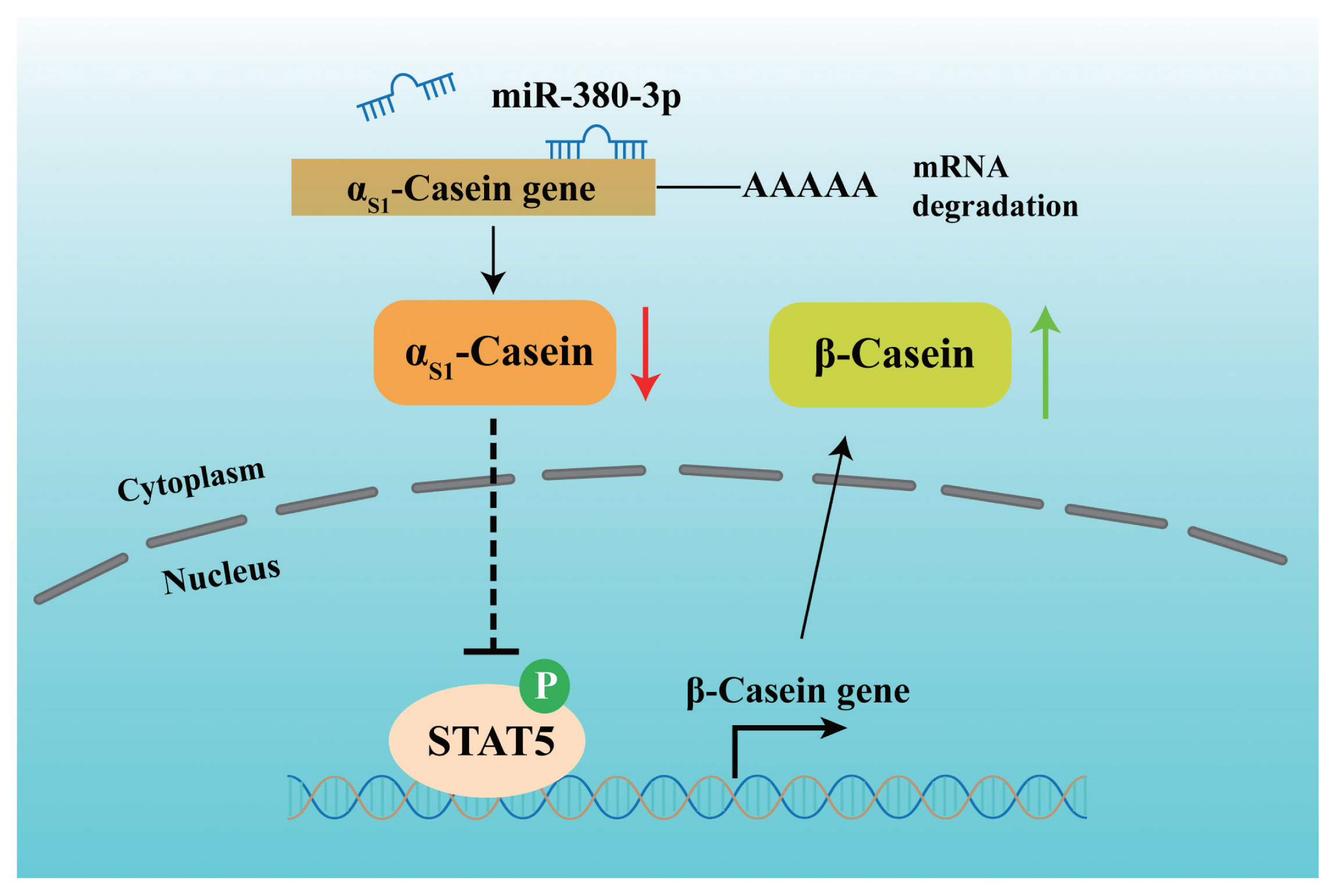
Further, we attempted to identify the binding of STAT5 to ╬▓-casein promoter by ChIP analysis. Compared with rabbit IgG control group, STAT5 forcefully bound to the specific site of ╬▓-casein promoter region in p-STAT5 group (p<0.01), but STAT5 inhibitor highly attenuated the combination of STAT5 and ╬▓-casein promoter region (p<0.01; Figure 5E). Subsequently, we discovered that the binding efficiency of STAT5 and ╬▓-casein promoter was significantly enhanced by miR-380-3p mimic treatment (p<0.05; Figure 5F). These data declared that STAT5 activated ╬▓-casein transcription and that the modulation of STAT5 on ╬▓-casein promoter was increased by miR-380-3p overexpression. Overall, our study revealed that miR-380-3p inhibits ╬▒S1-casein expression by targeting ╬▒S1-casein, and indirectly promotes ╬▓-casein transcription via ╬▒S1-casein/STAT5 axis in GMEC (Figure 6).
Understanding the regulatory mechanism of miRNA in physiological progresses could provide the theoretical basis for its application in animals. Previous studies have shown miRNAs modulate the synthesis process of milk proteins. miR-424-5p targeted 3ŌĆ▓UTR of ╬▓-casein gene by prolactin receptor signaling pathway in murine mammary glands [10]. miR-2904 positively regulated milk protein synthesis via Janus kinase 2 (JAK2)/STAT5 signaling pathway by the functional network analysis of Chinese Holstein cows [11]. In bovine mammary epithelial cells (BMEC), miR-139 inhibited ╬▓-casein expression through targeting growth hormone receptor and insulin-like growth factor receptor [12]. miR-27a negatively regulated milk protein and lactose contents through targeting mitogen-activated protein kinase 14 that had a positive effect on milk protein expression in sheep [13]. In GMEC, miR-8516 promoted the secretion of ╬▒S1-casein and ╬▓-casein by activating the activity of JAK2/STAT5 [14]. Although these studies indicate the miRNA regulation of milk protein synthesis, research on miRNA-targeted ╬▒S1-casein expression are still insufficient. According to the TargetScan software prediction, miR-380-3p targets 3ŌĆ▓UTR region of goat ╬▒S1-casein gene. A dual-luciferase experiment was carried out to identify the targeted relationship of miR-380-3p to ╬▒S1-casein gene. More importantly, miR-380-3p decreased the expression of ╬▒S1-casein gene and ╬▒S1-casein in dairy goats, which contributed to low allergy potential in goat milk.
miR-380-3p is involved in multiple biological progresses, such as cell proliferation, apoptosis, inflammation and melanogenesis. miR-380-3p targeted sex determining region Y-box 6 to regulate melanogenesis by affecting beta-catenin and microphthalmia-associated transcription factor expression in melanocytes [15]. High expression of miR-380-3p increased cell inflammation by promoting tumor necrosis factor-alpha, interleukin-1 beta, and interleukin-18 levels in spinal cord jury model rats [16]. Overexpression of miR-380-3p activated inflammation, oxidative stress, and apoptosis by targeting suppressor of cytokine signaling 6 in retinal epithelial cells with diabetic retinopathy [17]. In mouse neuroblastoma cells, miR-380-3p inhibited cell proliferation and increased cell apoptosis rate through blocking the translation of transcription factor specificity protein-3 [18]. On the contrary, miR-380-3p, acted as an oncogenic miRNA, promoted cell proliferation, epithelial-mesenchymal transition, and tumorigenesis in pancreatic cancer cells [19]. Despite the potent features of miR-380-3p, its regulatory role in milk protein expression is still unknown. In our research on dairy goats, we uncovered that miR-380-3p decreased ╬▒S1-casein expression and enhanced ╬▓-casein abundance by targeting 3ŌĆ▓UTR of ╬▒S1-casein gene. We expected that there was a correlation between ╬▒S1-casein and ╬▓-casein abundance regulated by miR-380-3p, which should be investigated in more detail in GMEC.
Multiple studies have shown the association between the expression of different milk protein components. In mouse mammary epithelial cells, lipoteichoic acid treatment increased the intracellular and secreted ╬▒S1-casein accompanied by a decrease in ╬▓-casein content [20]. Compared with wild-type dairy cows, the contents of ╬▒S1-casein, ╬▓-casein, ╬║-casein and ╬▒-lactalbumin were all increased in ╬▓-lactoglobulin silenced cows [21]. On the contrast, the contents of ╬▒-casein and ╬▒-lactalbumin in milk from ╬▓-casein and ╬║-casein transgenic cows were lower than non-transgenic cows [22]. There was a negative correlation between ╬▒S1-casein and ╬▓-casein gene expression in BMEC treated with camellia seed oil [23]. In mammary gland of lactation goats, ╬▒S1-casein gene expression was negatively correlated with ╬▓-lactoglobulin mRNA level [24]. More importantly, our previous study found that ╬▒S1-casein gene negatively regulated STAT5a phosphorylation and ╬▓-casein expression in GMEC [8]. In this research on GMEC, we carried out rescue assays to identify whether miR-380-3p up-regulated p-STAT5a and ╬▓-casein abundance by targeting ╬▒S1-casein gene. miR-380-3p mimic rescued the inhibition of ╬▒S1-casein gene on p-STAT5a and ╬▓-casein expression, while miR-380-3p inhibitor moderated the impact of ╬▒S1-casein gene knockdown on p-STAT5a and ╬▓-casein abundance. These rescue assays reveal that miR-380-3p indirectly increases STAT5 activity and ╬▓-casein expression by inhibiting ╬▒S1-casein expression. The present results are supported by our previous study that ╬▒S1-casein downregulated STAT5 activity and ╬▓-casein expression.
Previous studies have shown STAT5 is a primary transcriptional factor for milk production in mammals. STAT5 binds to milk protein gene promoters by identifying the specific binding motif. In STAT5 knockout mouse model, there was a significant reduction in the content of ╬▒-lactalbumin and ╬▓-lactoglobulin in milk [25]. Curcumin and basolateral lactose exposure suppressed intracellular and secreted casein production by STAT5 inactivation in mouse mammary epithelial cells [26]. In BMEC, beta-sitosterol promoted ╬▓-casein synthesis through the activation of STAT5 [27]; biochanin A declined ╬▓-casein synthesis and secretion by inhibiting STAT5 activity [28]. Milk protein expression was markedly decreased when STAT5a activity was blocked by suppressor of cytokine signaling 3 in dairy cows [29]. Further, phosphorylated STAT5a bound to ╬▓-casein promoter region and increased ╬▓-casein synthesis in immortalized GMEC [30]. Consistent with previous studies, luciferase experiments have shown that phosphorylated STAT5 promoted ╬▓-casein gene transcription as well in the present study. More importantly, ChIP experiments were carried out to demonstrate the direct binding of STAT5 to ╬▓-casein gene promoter, which could be activated by miR-380-3p mimic. Therefore, miR-380-3p indirectly promotes ╬▓-casein transcription through ╬▒S1-casein/STAT5a regulation axis. Additionally, ╬▒S1-casein gene could be targeted by miR-380-3p both in dairy goats and cows via TargetScan database prediction. In this study, we verified that miR-380-3p targets ╬▒S1-casein gene and promotes STAT5 activity and ╬▓-casein expression by inhibiting ╬▒S1-casein expression in dairy goats. However, whether miR-380-3p could target ╬▒S1-casein gene and play similar regulatory roles needs to be further studied in dairy cows.
miR-380-3p targets 3ŌĆ▓UTR of goat ╬▒S1-casein gene and inhibits ╬▒S1-casein expression which is associated with higher rates of milk allergy for humans. miR-380-3p promotes ╬▓-casein expression through targeting ╬▒S1-casein gene. Moreover, miR-380-3p increases ╬▓-casein transcription by the binding of STAT5 to ╬▓-casein gene promoter region. In summary, our study is helpful to conclude the regulatory role of miR-380-3p on milk protein synthesis and supplies a novel basis for declining the potential of milk allergy through regulating miR-380-3p in ruminants.
Notes
SUPPLEMENTARY MATERIAL
Supplementary file is available from: https://doi.org/10.5713/ab.23.0007
Supplementary Table S1. Primers used for amplification of ╬▒S1-casein and ╬▓-casein gene
ab-23-0007-Supplementary-Table-1.pdf
Supplementary Table S2. Sequences of siRNA targeting goat ╬▒S1-casein gene
ab-23-0007-Supplementary-Table-2.pdf
Supplementary Figure S1. The image of the full western blot of Figure 2C. Expression of ╬▒S1-casein in cells treated with miR-380-3p mimic (50 nM) or miR-380-3p inhibitor (100 nM) for 48 h.
ab-23-0007-Supplementary-Fig-1.pdf
Supplementary Figure S2. The image of the full western blot of Figure 3C. Expression of p-STAT5a and ╬▓-casein in cells treated with miR-380-3p mimic (50 nM) or miR-380-3p inhibitor (100 nM) for 48 h.
ab-23-0007-Supplementary-Fig-2.pdf
Supplementary Figure S3. The image of the full western blot of Figure 4B. Expression of ╬▒S1-casein, p-STAT5a and ╬▓-casein in cells co-treated with pcDNA3.1-╬▒S1-casein and miR-380-3p mimic (50 nM) for 48 h.
ab-23-0007-Supplementary-Fig-3.pdf
Supplementary Figure S4. The image of the full western blot of Figure 4D. Expression of ╬▒S1-casein, p-STAT5a and ╬▓-casein in cells co-treated with si-╬▒S1-Casein (100 nM) and miR-380-3p inhibitor (100 nM) for 48 h.
ab-23-0007-Supplementary-Fig-4.pdf
Figure┬Ā1
miR-380-3p targets 3ŌĆ▓UTR of ╬▒S1-casein gene. (A) ╬▒S1-Casein gene and miR-380-3p abundance in goat mammary gland of different lactation stages. Mammary gland samples were obtained from Xinong Saanen dairy goats (n = 5) during early, peak, middle and end lactation stages. Values were normalized to early lactation stage. (B) Predicted binding site of miR-380-3p in ╬▒S1-casein 3ŌĆ▓UTR and construction of psiCHECK-2 dual-luciferase reporter vectors. psi-WT-╬▒S1-casein represents the wildtype sequence vector; psi-MUT-╬▒S1-casein represents the mutation (red bases) vector. (C) miR-380-3p expression was measured by qRT-PCR. GMEC were treated with mimic NC or miR-380-3p mimic (50 nM) for 48 h. (D) 3ŌĆ▓UTR activity of ╬▒S1-casein was measured by luciferase assay. Cells were co-treated with dual-luciferase vector and miR-380-3p mimic (50 nM) for 48 h. Data are presented as mean┬▒standard error of the mean. UTR, untranslated region; qRT-PCR, quantitative real-time polymerase chain reaction; GMEC, goat mammary epithelial cells; NC, negative control. aŌĆōc Different superscripts represent significant differences among ╬▒S1-casein or miR-380-3p levels (p<0.05). * p<0.05 and ** p<0.01 compared with control.
Figure┬Ā2
miR-380-3p inhibits ╬▒S1-casein expression. (A) Expression of ╬▒S1-casein gene in cells treated with miR-380-3p mimic (50 nM) for 48 h. (B) Expression of ╬▒S1-casein gene in cells treated with miR-380-3p inhibitor (100 nM) for 48 h. (C) Expression of ╬▒S1-casein in cells treated with miR-380-3p mimic (50 nM) or miR-380-3p inhibitor (100 nM) for 48 h. Relative ╬▒S1-casein expression was normalized by comparison with ╬▓-actin. Data are presented as mean┬▒standard error of the mean. * p<0.05 and ** p<0.01 compared with control.

Figure┬Ā3
miR-380-3p increases STAT5a activity and ╬▓-casein abundance. (A) Expression of ╬▒S2-casein, ╬▓-casein and ╬║-casein gene in cells treated with miR-380-3p mimic (50 nM) for 48 h. (B) Expression of ╬▒S2-casein, ╬▓-casein and ╬║-casein gene in cells treated with miR-380-3p inhibitor (100 nM) for 48 h. (C) Expression of p-STAT5a and ╬▓-casein in cells treated with miR-380-3p mimic (50 nM) or miR-380-3p inhibitor (100 nM) for 48 h. Relative ╬▓-casein expression was normalized by comparison with ╬▓-actin. Relative p-STAT5a expression was normalized to STAT5a. STAT5a, signal transducer and activator of transcription 5a. Data are presented as mean┬▒standard error of the mean. * p<0.05 and ** p<0.01 compared with control.

Figure┬Ā4
miR-380-3p improves ╬▓-casein expression by targeting ╬▒S1-casein gene. (A, B) Expression of ╬▒S1-casein, p-STAT5a and ╬▓-casein in cells co-treated with pcDNA3.1-╬▒S1-casein and miR-380-3p mimic (50 nM) for 48 h. (C, D) Expression of ╬▒S1-casein, p-STAT5a and ╬▓-casein in cells co-treated with si-╬▒S1-Casein (100 nM) and miR-380-3p inhibitor (100 nM) for 48 h. Relative expression of ╬▒S1-casein and ╬▓-casein was normalized by comparison with ╬▓-actin. Relative p-STAT5a expression was normalized to STAT5a. STAT5a, signal transducer and activator of transcription 5a. Data are presented as mean┬▒standard error of the mean. aŌĆōc Different superscripts represent significant differences among groups (p<0.05).

Figure┬Ā5
miR-380-3p activates ╬▓-casein transcription via regulating STAT5. (A) Conserved binding motif of transcription factor STAT5. (B) Binding site of STAT5 in ╬▓-casein promoter region and construction of pGL3 luciferase reporter vectors. pGL3-WT-╬▓-casein represents the wildtype ╬▓-casein promoter vector; pGL3-MUT-╬▓-Casein represents the mutation (red bases) vector on STAT5 binding sequence. (C) Promoter activity of ╬▓-casein was measured by luciferase assay. Cells were treated with STAT5-IN-1 (50 ╬╝M) followed by pGL3-WT-╬▓-Casein and miR-380-3p mimic (50 nM) co-treatment for 48 h. (D) Relative luciferase activity in cells co-treated with pGL3-╬▓-Casein and miR-380-3p mimic (50 nM) for 48 h. (E) ChIP assay of the STAT5 binding site to ╬▓-casein gene promoter by qRT-PCR. Cells were treated with STAT5-IN-1 (50 ╬╝M) for 48 h. (F) ChIP assay in cells treated with miR-380-3p mimic (50 nM) for 48 h. STAT5a, signal transducer and activator of transcription 5a; qRT-PCR, quantitative real-time polymerase chain reaction. Data are presented as mean┬▒standard error of the mean. * p<0.05 and ** p<0.01 compared with control.
Figure┬Ā6
The proposed model regarding the role and regulation of miR-380-3p in goat mammary epithelial cells. miR-380-3p inhibits ╬▒S1-casein expression by targeting 3ŌĆ▓UTR of ╬▒S1-casein gene. Then, miR-380-3p increases ╬▓-casein expression through ╬▒S1-casein/STAT5 axis. Moreover, miR-380-3p promotes ╬▓-casein transcription by the binding of STAT5 to ╬▓-casein gene promoter region. STAT5a, signal transducer and activator of transcription 5a.
Table┬Ā1
Primers used for quantitative real-time polymerase chain reaction of genes and promoter region
REFERENCES
1. Hochwallner H, Schulmeister U, Swoboda I, et al. Infant milk formulas differ regarding their allergenic activity and induction of T-cell and cytokine responses. Allergy 2017; 72:416ŌĆō24.
https://doi.org/10.1111/all.12992



2. Hochwallner H, Schulmeister U, Swoboda I, Spitzauer S, Valenta R. CowŌĆÖs milk allergy: From allergens to new forms of diagnosis, therapy and prevention. Methods 2014; 66:22ŌĆō33.
https://doi.org/10.1016/j.ymeth.2013.08.005



3. Caubet JC, Lin J, Ahrens B, et al. Natural tolerance development in cowŌĆÖs milk allergic children: IgE and IgG4 epitope binding. Allergy 2017; 72:1677ŌĆō85.
https://doi.org/10.1111/all.13167


4. Carillier-Jacquin C, Larroque H, Robert-Granie C. Including a(s1) casein gene information in genomic evaluations of French dairy goats. Genet Sel Evol 2016; 48:54
https://doi.org/10.1186/s12711-016-0233-x



5. Wang Q, Ren N, Cai Z, et al. Paraquat and MPTP induce neurodegeneration and alteration in the expression profile of microRNAs: the role of transcription factor Nrf2. NPJ Parkinsons Dis 2017; 3:31
https://doi.org/10.1038/s41531-017-0033-1



6. Li S, Wu D, Jia H, Zhang Z. Long non-coding RNA LRRC75A-AS1 facilitates triple negative breast cancer cell proliferation and invasion via functioning as a ceRNA to modulate BAALC. Cell Death Dis 2020; 11:643
https://doi.org/10.1038/s41419-020-02821-2



7. Holliday H, Yang J, Dodson E, et al. miR-99b-5p, miR-380-3p, and miR-485-3p are novel chemosensitizing miRNAs in high-risk neuroblastoma. Mol Ther 2022; 30:1119ŌĆō34.
https://doi.org/10.1016/j.ymthe.2022.01.004



8. Song N, Chen Y, Luo J, et al. Negative regulation of alpha(S1)-casein (CSN1S1) improves beta-casein content and reduces allergy potential in goat milk. J Dairy Sci 2020; 103:9561ŌĆō72.
https://doi.org/10.3168/jds.2020-18595


9. Kung MH, Lee YJ, Hsu JT, Huang MC, Ju YT. A functional study of proximal goat beta-casein promoter and intron 1 in immortalized goat mammary epithelial cells. J Dairy Sci 2015; 98:3859ŌĆō75.
https://doi.org/10.3168/jds.2014-9054


10. Wang W, Zang X, Liu Y, et al. Dynamic miRNA landscape links mammary gland development to the regulation of milk protein expression in mice. Animals 2022; 12:727
https://doi.org/10.3390/ani12060727



11. Cai W, Li C, Li J, Song J, Zhang S. Integrated small RNA sequencing, transcriptome and GWAS data reveal microRNA regulation in response to milk protein traits in Chinese Holstein cattle. Front Genet 2021; 12:726706
https://doi.org/10.3389/fgene.2021.726706



12. Cui Y, Sun X, Jin L, et al. MiR-139 suppresses beta-casein synthesis and proliferation in bovine mammary epithelial cells by targeting the GHR and IGF1R signaling pathways. BMC Vet Res 2017; 13:350
https://doi.org/10.1186/s12917-017-1267-1



13. Duman E, Ozmen O, Kul S. Oar-miR-16b and oar-miR-27a: negatively correlated with milk yield and milk protein in sheep. Anim Biotechnol 2022; 33:1466ŌĆō79.
https://doi.org/10.1080/10495398.2021.1908317


14. Zhang Y, Wu Q, Liu J, An X, Cao B. Circ-140/chi-miR-8516/STC1-MMP1 regulates alpha s1-/beta-casein secretion and lipid formation in goat mammary epithelial cells. Genes 2021; 12:671
https://doi.org/10.3390/genes12050671



15. Liu X, Du B, Zhang P, et al. miR-380-3p regulates melanogenesis by targeting SOX6 in melanocytes from alpacas (Vicugna pacos). BMC Genomics 2019; 20:962
https://doi.org/10.1186/s12864-019-6343-4



16. Li X, Lou X, Xu S, Du J, Wu J. Hypoxia inducible factor-1 (HIF-1 alpha) reduced inflammation in spinal cord injury via miR-380-3p/NLRP3 by Circ 0001723. Biol Res 2020; 53:35
https://doi.org/10.1186/s40659-020-00302-6



17. Luo R, Li L, Xiao F, Fu J. LncRNA FLG-AS1 mitigates diabetic retinopathy by regulating retinal epithelial cell inflammation, oxidative stress, and apoptosis via miR-380-3p/SOCS6 axis. Inflammation 2022; 45:1936ŌĆō49.
https://doi.org/10.1007/s10753-022-01665-6


18. Cai Z, Zheng F, Ding Y, et al. Nrf2-regulated miR-380-3p blocks the translation of Sp3 protein and its mediation of paraquat-induced toxicity in mouse neuroblastoma N2a cells. Toxicol Sci 2019; 171:515ŌĆō29.
https://doi.org/10.1093/toxsci/kfz162



19. Jiang Z, Song X, Wei Y, Li Y, Kong D, Sun J. N(6)-methyladenosine-mediated miR-380-3p maturation and upregulation promotes cancer aggressiveness in pancreatic cancer. Bioengineered 2022; 13:14460ŌĆō71.
https://doi.org/10.1080/21655979.2022.2088497



20. Kobayashi K, Omatsu N, Han L, Shan-Ni L, Nishimura T. Early effects of lipoteichoic acid from Staphylococcus aureus on milk production-related signaling pathways in mouse mammary epithelial cells. Exp Cell Res 2022; 420:113352
https://doi.org/10.1016/j.yexcr.2022.113352


21. Jabed A, Wagner S, McCracken J, Wells DN, Laible G. Targeted microRNA expression in dairy cattle directs production of beta-lactoglobulin-free, high-casein milk. Proc Natl Acad Sci USA 2012; 109:16811ŌĆō6.
https://doi.org/10.1073/pnas.1210057109



22. Laible G, Smolenski G, Wheeler T, Brophy B. Increased gene dosage for beta- and kappa-casein in transgenic cattle improves milk composition through complex effects. Sci Rep 2016; 6:37607
https://doi.org/10.1038/srep37607



23. Zhong W, Shen J, Liao X, et al. Camellia (Camellia oleifera Abel.) seed oil promotes milk fat and protein synthesis-related gene expression in bovine mammary epithelial cells. Food Sci Nutr 2020; 8:419ŌĆō27.
https://doi.org/10.1002/fsn3.1326



24. Morammazi S, Masoudi AA, Torshizi RV, Pakdel A. Differential expression of the Alpha S1 casein and Beta-lactoglobulin genes in different physiological stages of the Adani goats mammary glands. Iran J Biotechnol 2016; 14:278ŌĆō85.
https://doi.org/10.15171/ijb.1171



25. Reichenstein M, Rauner G, Barash T. Conditional repression of STAT5 expression during lactation reveals its exclusive roles in mammary gland morphology, milk-protein gene expression, and neonate growth. Mol Reprod Dev 2011; 78:585ŌĆō96.
https://doi.org/10.1002/mrd.21345


26. Kobayashi K, Tsugami Y, Suzuki N, Suzuki T, Nishimura T. Lactose on the basolateral side of mammary epithelial cells inhibits milk production concomitantly with signal transducer and activator of transcription 5 inactivation. Cell Tissue Res 2022; 389:501ŌĆō15.
https://doi.org/10.1007/s00441-022-03651-8


27. Liu X, Shen J, Zong J, Liu J, Jin Y. Beta-sitosterol promotes milk protein and fat syntheses-related genes in bovine mammary epithelial cells. Animals 2021; 11:3238
https://doi.org/10.3390/ani11113238



28. Tsugami Y, Wakasa H, Kawahara M, Nishimura T, Kobayashi K. Isoflavones and their metabolites influence the milk production ability of bovine mammary epithelial cells in a type-specific manner. J Anim Sci 2022; 93:e13720
https://doi.org/10.1111/asj.13720

29. Khan MZ, Khan A, Xiao J, et al. Role of the JAK-STAT pathway in bovine mastitis and milk production. Animals 2020; 10:2107
https://doi.org/10.3390/ani10112107



30. Ghosh A, Radhakrishnan R. Time-dependent antagonist-agonist switching in receptor tyrosine kinase-mediated signaling. BMC Bioinformatics 2019; 20:242
https://doi.org/10.1186/s12859-019-2816-3



- TOOLS
-
METRICS

-
- 1 Crossref
- 0 Scopus
- 2,691 View
- 147 Download
- Related articles






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement1
Supplement1 Print
Print




